
Researchers develop AI-based tool paving the way for personalized cancer treatments
In the ongoing fight against cancer, scientists around the globe are exploring innovative approaches to unlock the mysteries of the human immune system — the complex network of organs, cells and proteins that defends the body against disease.
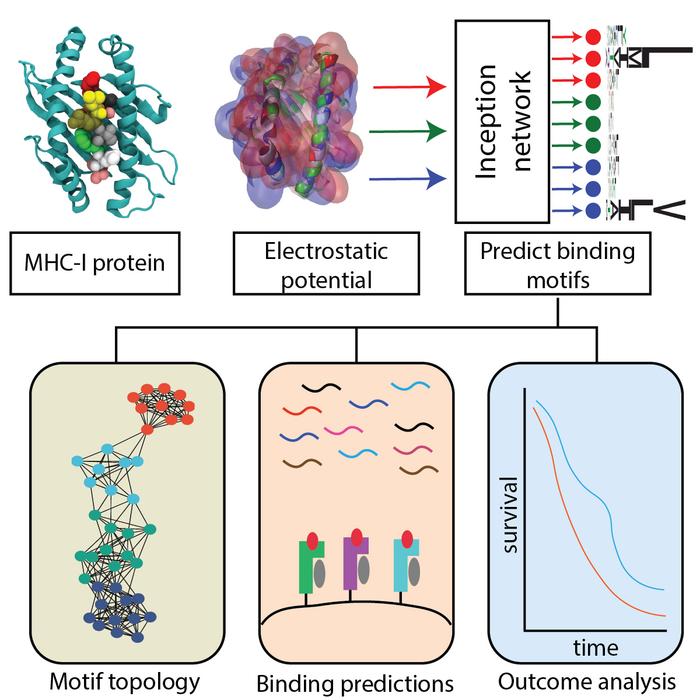
Credit: Arizona State University
In the ongoing fight against cancer, scientists around the globe are exploring innovative approaches to unlock the mysteries of the human immune system — the complex network of organs, cells and proteins that defends the body against disease.
A team led by Arizona State University scientists have developed an AI-based learning tool called HLA Inception that’s uncovered new information about how an individual person’s immune system responds to foreign cells.
Focusing on a group of proteins called Major Histocompatibility Complex-1(MHC-1), the AI-based tool, in seconds, can classify the specific group of proteins unique for an individual and predict whether a person’s immune defenses may recognize pieces of threatening viruses and cancers.
“We are able to make predictions on pathological outcomes of patients, such as the survival against certain cancer medicines, based on the molecular details that a human is born with,” said Abhishek Singharoy, assistant professor in ASU’s School of Molecular Sciences who led the study. “Now with this tool, something that was taking days only takes seconds.”
Understanding this individualized molecular interaction information holds tremendous promise for creating new personalized cancer medicines with the potential to transform patient care.
The work was published in the journal of Cell Systems on March 29.
Unraveling the complexities MHC-1 Protein Preferences
Within the human body, MHC-1 proteins act as guards on the surface of our cells alerting the immune system to foreign invaders. They grab pieces of foreign proteins, or peptides, inside cells and present them to the immune system for recognition and attack.
Each person’s MHC-1 proteins have specific preferences for the types of protein fragments they interact with and being able to predict which peptides will bind effectively to which MHC-I molecules is crucial for further understanding the mechanisms of how our immune system works and for developing new more advanced cancer vaccines.
However, prediction is challenging.
There are thousands of different versions of MHC-I molecules in the human population, making it hard to create a universal prediction model.
Analyzing nearly 6,000 MHC-1 complexes, the research team discovered patterns that can identify these preferences and predict immune responses across a broad range of human populations.
The HLA Inception tool, powered by AI and machine learning, uses the varied charges on the surface of the proteins, also known as the electrostatic signatures, to classify them into 11 different types.
This information can then be used to predict whether the protein fragments, or peptides, MHC-1 are monitoring are self or foreign invaders (non-self).
The researchers also found that patients with a more diverse range of MHC-1 proteins, covering more of the 11 classes, had a higher chance of surviving certain cancer therapies.
“The continued integration of machine learning in health care will help de-risk and personalize treatments,” said Eric Wilson, an author on the paper, ASU alumnus and currently a postdoctoral fellow at Icahn School of Medicine at Mount Sinai. “Machine learning and AI can improve the accessibility of new treatments to a broader cohort of patients by negating the need for costly experiments to determine candidacy.”
Committed to advancing scientific progress in the field, the researchers have made HLA-Inception freely available for academic use, laying the foundation for widespread collaboration and innovation in the field of immunotherapy.
“I am excited to use these tools for the development of better cancer therapeutic vaccines and immunotherapies,” said Karen Anderson, a co-author on the paper and professor in ASU’s School of Life Sciences. “This is the ultimate approach for precision medicine. Next-generation immunotherapies will be highly precise and tailored based on an individual’s MHC molecules.”
The researchers envision this work contributing to advancements in healthcare, particularly for tailoring treatments to individual patients.
“This is impactful research with ramifications beyond the limits of academia,” said Singharoy, who is also a researcher with the ASU Biodesign Center for Applied Structural Discovery. “Our technique now is the fastest out there.”
Additional researchers involved in the study include John Kevin Cava, Arizona State University’s School of Computing and Augmented Intelligence; Diego Chowell, The Precision Immunology Institute, Icahn School of Medicine at Mount Sinai; and Remya Raja, Kiran K. Mangalaparthi, Akhilesh Pandey and Marion Curtis, Mayo Clinic.
Journal
Cell Systems
DOI
10.1016/j.cels.2024.03.001
Method of Research
Data/statistical analysis
Subject of Research
Cells
Article Title
The Electrostatic Landscape of MHC-peptide Binding Revealed Using Inception Networks
Article Publication Date
29-Mar-2024