Exploring the role of seven key genes in breast cancer: insights from in silico and in vitro analyses
Breast cancer remains a significant global health concern, warranting further exploration into its genetic basis and potential therapeutic targets. This study aimed to elucidate the genetic associations of seven pivotal genes with breast cancer and discern their potential role in disease prognosis.
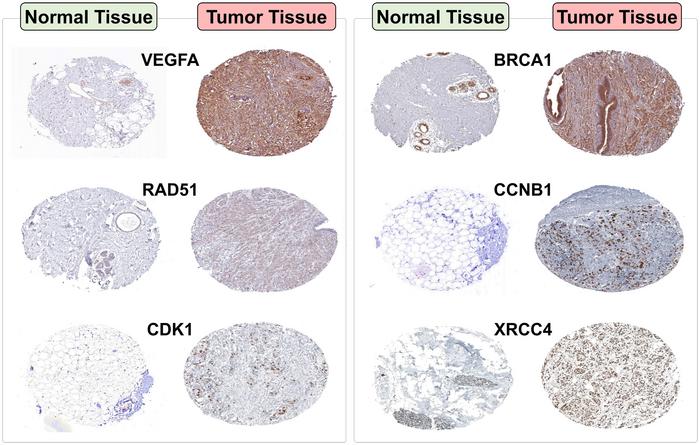
Credit: Rakesh M. Rawal, Devashish Mehta
Background and objectives
Breast cancer remains a significant global health concern, warranting further exploration into its genetic basis and potential therapeutic targets. This study aimed to elucidate the genetic associations of seven pivotal genes with breast cancer and discern their potential role in disease prognosis.
Methods
The genes VEGFA, BRCA1, RAD51, CCNB1, CHEK1, CDK1, and XRCC4 were curated from over 30 articles. Their association with breast cancer was analyzed using both in silico and in vitro techniques. The in silico assessment involved constructing a protein-protein interaction network, accompanied by Gene Ontology and pathway enrichment analysis. Further, survival and expression analysis were conducted using Kaplan-Meier Plotter and the UALCAN database respectively. At the protein level, expression was observed using the Human Protein Atlas database. The in vitro validation involved analyzing mRNA expression levels in 10 breast cancer tissue samples.
Results
The study revealed that all seven genes are significantly upregulated in breast cancer tissues compared to normal tissues, highlighting their critical role in tumor development and progression. Protein-protein interaction analysis confirmed their central involvement in vital biological processes related to breast cancer. Survival analysis showed that high expressions of these genes are associated with poorer patient prognosis, with hazard ratios indicating their potential as prognostic markers. In vitro validation further supported their overexpression in breast cancer, suggesting their importance in molecular landscape of the disease and their value as targets for therapeutic intervention.
Conclusions
This study contributes to the understanding of the genetic landscape of BC and underscores the potential of seven genes studied as targets for therapeutic intervention. The findings emphasize the importance of these genes in the pathology of BC and pave the way for personalized medicine. Future research should focus on exploring the therapeutic potential of these genes and developing targeted therapies for BC. This could involve functional studies to elucidate the precise roles of these genes in BC and preclinical studies to evaluate the efficacy of targeted therapies. Ultimately, these efforts could lead to improved treatment strategies and outcomes for patients with BC.
Full text
https://www.xiahepublishing.com/1555-3884/GE-2023-00094
The study was recently published in the Gene Expression.
Gene Expression (GE) is an open-access journal. It was launched in 1991 by Chicago Medical School Press, and transferred to Cognizant Communication Corporation in 1994. From August 2022, GE is published by Xia & He Publishing Inc.
GE publishes peer-reviewed and high-quality original articles, reviews, editorials, commentaries, and opinions on its primary research topics including cell biology, molecular biology, genes, and genetics, especially on the cellular and molecular mechanisms of human diseases.
GE has been indexed in Medline (1991-2021), Scopus, Biological Abstracts, Biosis Previews, ProQuest, etc.
Follow us on X: https://twitter.com/xiahepublishing
Follow us on LinkedIn: https://www.linkedin.com/company/xia&he-publishing-inc/
Journal
Gene Expression
DOI
10.14218/GE.2023.00094
Article Title
Exploring the Role of Seven Key Genes in Breast Cancer: Insights from In Silico and In Vitro Analyses
Article Publication Date
2-Jan-2024